Basics of GIS mapping in R
Abstract
This webinar introduces the basics of GIS data manipulation and mapping in R with the packages sf and tmap.
We will use data on the retreat of North American glaciers as an example.
Video
Getting data
The datasets that we will use for this webinar are the Alaska as well as the Western Canada and USA subsets of the Randolph Glacier Inventory version 6.01 and the USGS time series of the named glaciers of Glacier National Park2 . These datasets can be downloaded as zip files from their respective websites.
For our basemap, we will use data from Natural Earth. This dataset can be accessed direction from within R thanks to the packages rnaturalearth (which provides the functions) and rnaturalearthdata (which provides the data).
-
RGI Consortium (2017). Randolph Glacier Inventory – A Dataset of Global Glacier Outlines: Version 6.0: Technical Report, Global Land Ice Measurements from Space, Colorado, USA. Digital Media. DOI: https://doi.org/10.7265/N5-RGI-60. ↩
-
Fagre, D.B., McKeon, L.A., Dick, K.A., and Fountain, A.G., 2017, Glacier margin time series (1966, 1998, 2005, 2015) of the named glaciers of Glacier National Park, MT, USA: U.S. Geological Survey data release, https://doi.org/10.5066/F7P26WB1. ↩
Loading and exploring data
First, let's load the necessary packages for this webinar:
library(sf)
library(tmap)
library(dplyr)
library(magrittr)
library(purrr)
library(rnaturalearth)
library(rnaturalearthdata)We will start by mapping the data contained in the Alaska and the Western Canada and USA subsets of the Randolph Glacier Inventory, i.e. all the glaciers of Western North America.
Data get imported and turned into sf objects by the function sf::st_read():
ak <- st_read("01_rgi60_Alaska")
wes <- st_read("02_rgi60_WesternCanadaUS")
ak
wes
str(ak)
str(wes)Note: 02_rgi60_WesternCanadaUS and 01_rgi60_Alaska are the 2 directories containing the downloaded and unzipped datasets. Make sure to use the absolute paths or the proper paths relative to your working directory (which can be obtained with getwd() and modified with setwd()).
sf objects are data.frame-like objects with a geometry list-column as their last column. That column is itself an object of class sfc (simple feature geometry list column).
Mapping with tmap
tmap follows a grammar of graphic similar to that of ggplot2: you first need to set a shape (a spatial data object) by passing an sf object to tm_shape(). Then you plot one or several layers with one of several tmap functions and you use the + sign between each element.
To see the available options, run:
?tmap-elementWe could thus plot the glaciers of Alaska with any of the options below:
tm_shape(ak) +
tm_borders()
tm_shape(ak) +
tm_fill()
tm_shape(ak) +
tm_polygons() # shows both borders and fill
Here, we will use tm_polygons() which combines tm_borders() and tm_fill().
Layout elements and attribute layers
A map without title, compass, or scale bars is not very useful though. We need to add layout elements and attribute layers to the map.
You can loop up the many arguments of the tmap functions in the help pages to see how you can customize your maps:
?tm_layout
?tm_compass
?tm_scale_barLet's now map the glaciers of Alaska:
tm_shape(ak) +
tm_polygons() +
tm_layout(
title = "Glaciers of Alaska",
title.position = c("center", "top"),
title.size = 1.1,
bg.color = "#fcfcfc",
inner.margins = c(0.06, 0.01, 0.09, 0.01),
outer.margins = 0,
frame.lwd = 0.2
) +
tm_compass(
type = "arrow",
position = c("right", "top"),
size = 1.2,
text.size = 0.6
) +
tm_scale_bar(
breaks = c(0, 500, 1000),
position = c("right", "BOTTOM")
)
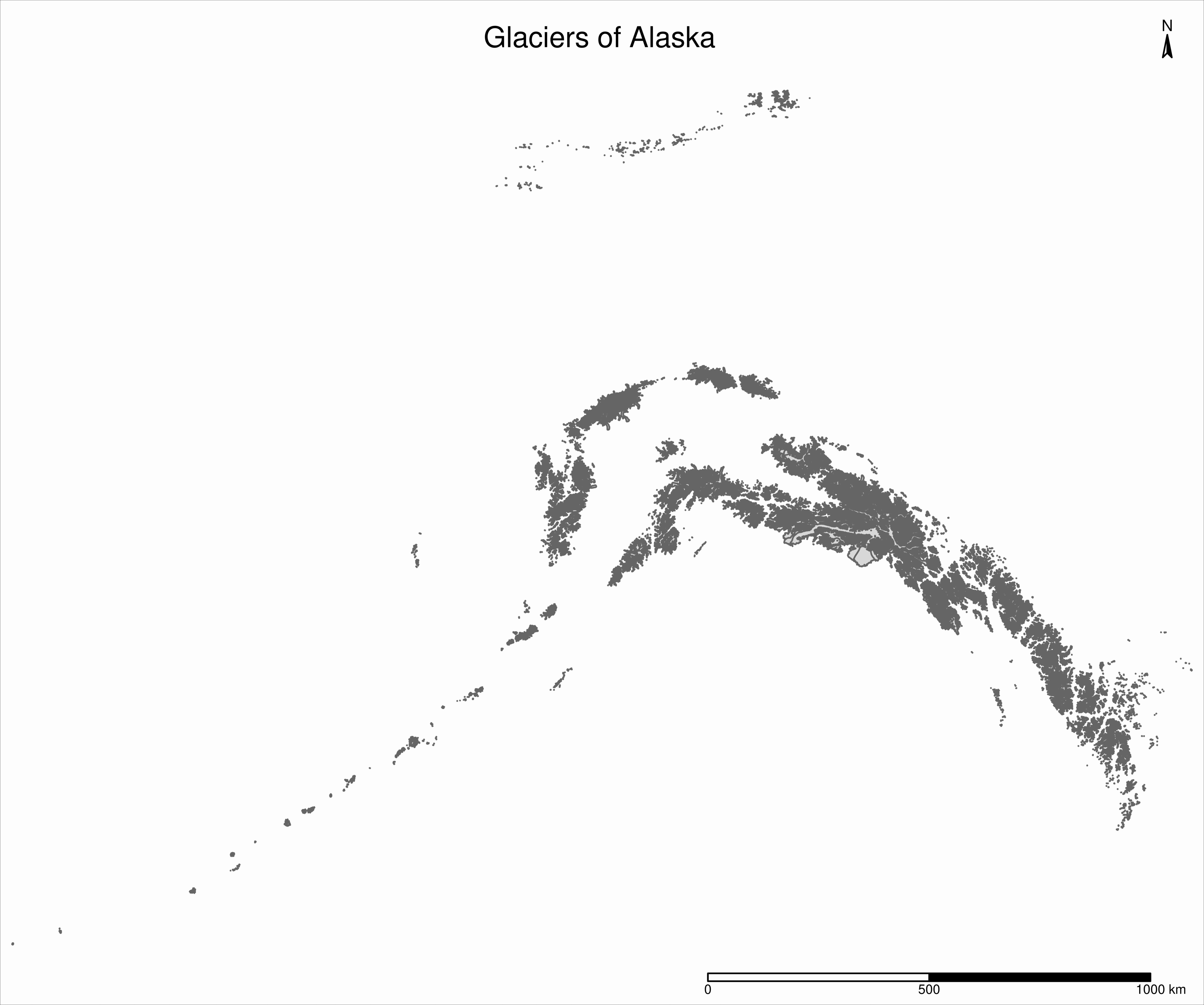
Union of bounding boxes
Now, if we want to plot all the glaciers of Western North America, we want to combine both sf objects in the same map. A map can contain multiple shapes: you only need to "add" a tm_shape and its element(s). Before doing so however, it is very important to ensure that they have the same coordinate reference system (CRS):
st_crs(ak)
st_crs(wes)
st_crs(ak) == st_crs(wes)They do, so we are good to go.
Note that, as with ggplot2 or GIS graphical user interfaces, the order matters since the layers stack up on top of each other.
tm_shape(ak) +
tm_polygons() +
tm_shape(wes) +
tm_polygons()If you run the code above however, you may be surprised that you are still only plotting the map of Alaska.
This is because each map comes with a spatial bounding box (bbox).
st_bbox(ak)
st_bbox(wes)
In the code above, the bbox is set by the first shape, i.e. our entire map uses the bbox of the Alaska sf object.
We first need to create a new bounding box encompassing both bounding boxes:
nwa_bbox <- st_bbox(ak) %>%
st_as_sfc() %>%
st_union(st_as_sfc(st_bbox(wes))) %>%
st_bbox()We can now plot the glaciers of Western North America:
tm_shape(ak, bbox = nwa_bbox) +
tm_polygons() +
tm_shape(wes) +
tm_polygons() +
tm_layout(
title = "Glaciers of Western North America",
title.position = c("center", "top"),
title.size = 1.1,
bg.color = "#fcfcfc",
inner.margins = c(0.06, 0.01, 0.09, 0.01),
outer.margins = 0,
frame.lwd = 0.2
) +
tm_compass(
type = "arrow",
position = c("right", "top"),
size = 1.2,
text.size = 0.6
) +
tm_scale_bar(
breaks = c(0, 1000, 2000),
position = c("right", "BOTTOM")
)
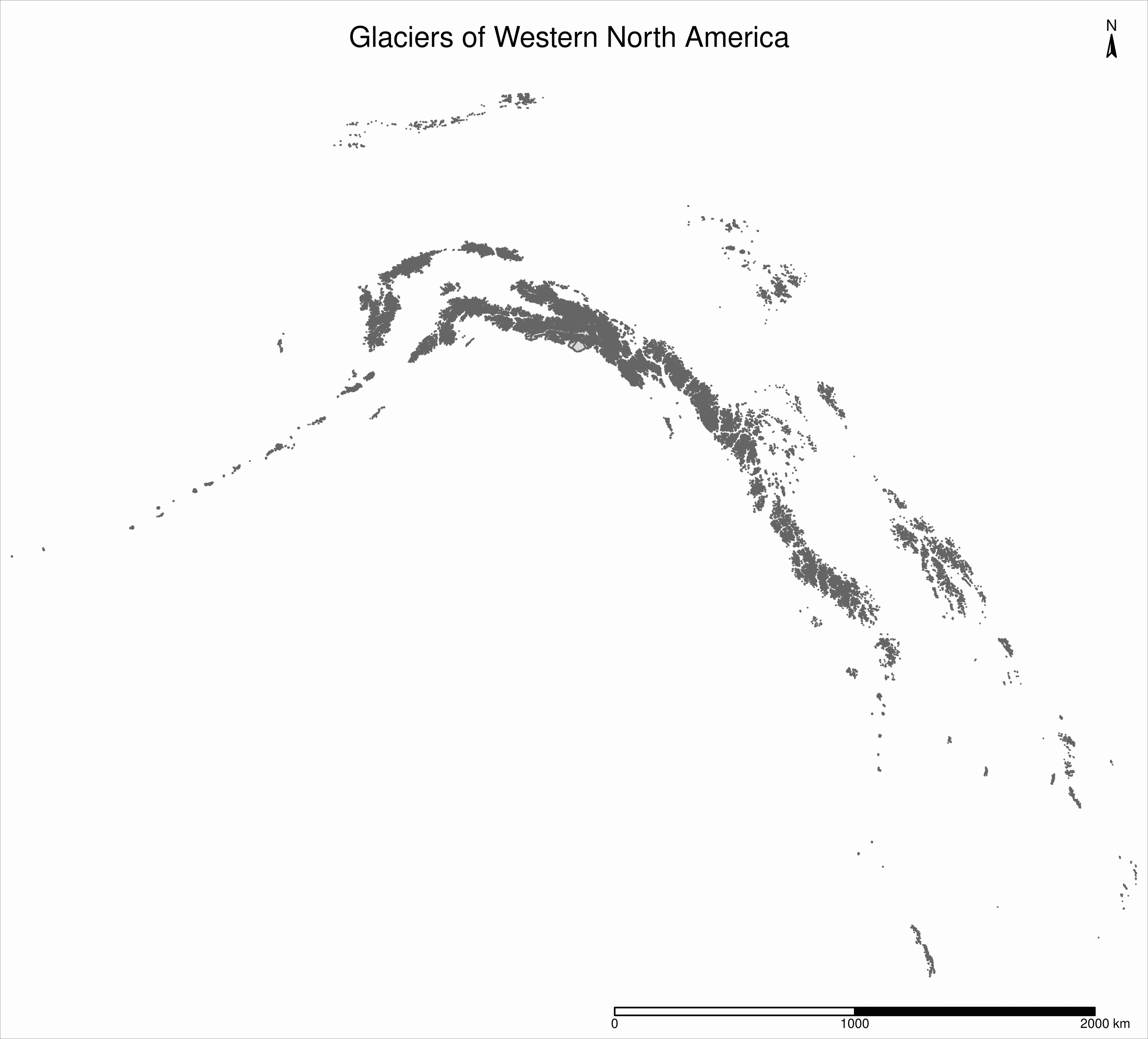
tmap maps with Leaflet
tmap has 2 modes: plot and view.
plot is the default mode for static maps that we have used so far.
view mode is an interactive viewing mode using leaflet in a browser. There, you can zoom in/out, select/deselect the different layers, and choose to display one of Esri.WorldGrayCanvas, OpenStreetMap, or Esri.WorldTopoMap basemaps.
You can toggle between the plot and view modes with ttm().
You can also switch to either mode with tmap_mode("view") and tmap_mode("plot").
tmap_mode("view")
tmap_last()
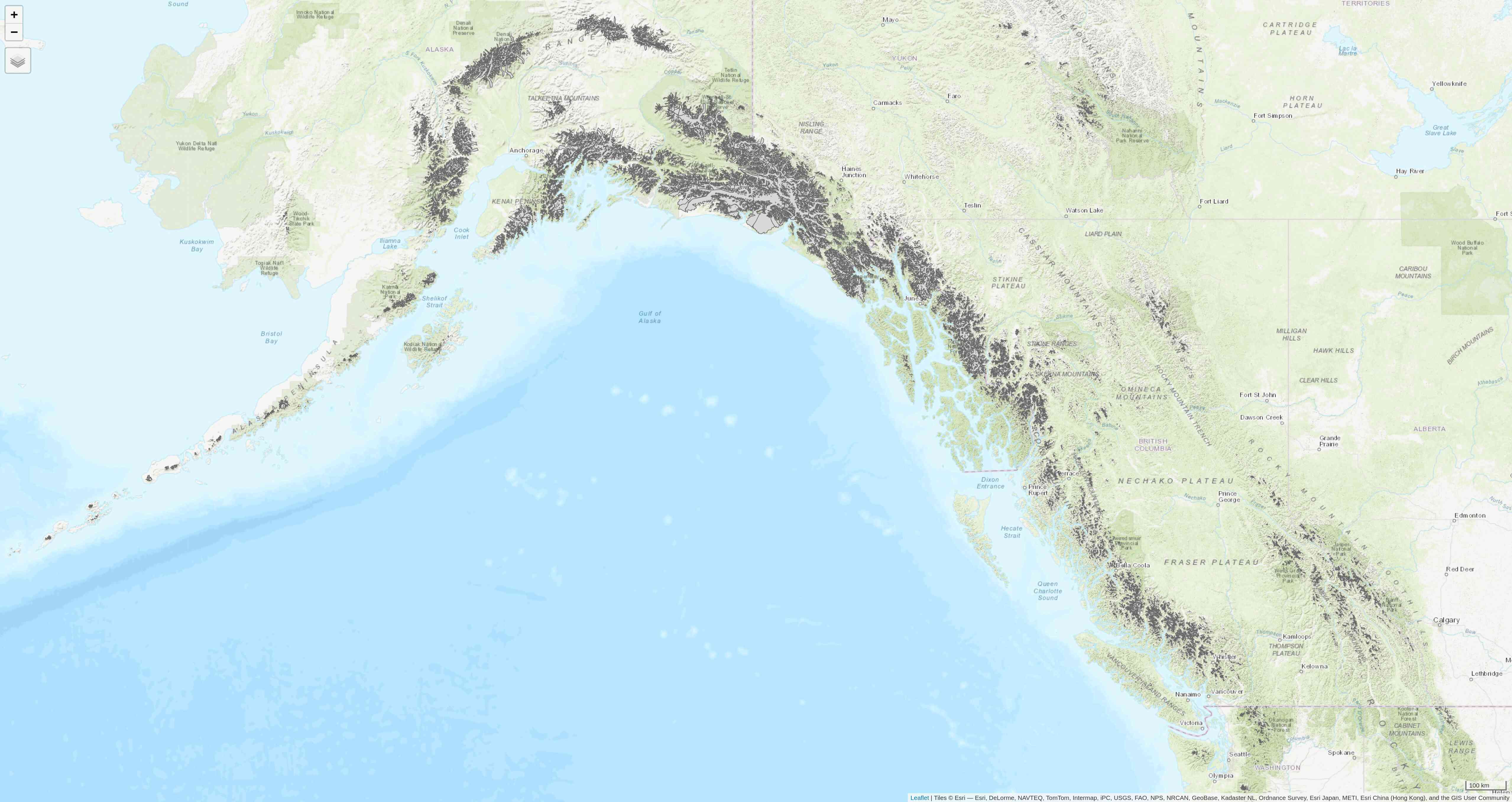
Afterwards, if you want to create new static plots, don't forget to get back to plot mode with either of:
ttmp()
tmap_mode("plot")Maps based on an attribute variable
What is interesting about glacier maps is to see their evolution through time as glaciers retreat due to climate change. While the Randolph Glacier Inventory (RGI) has an amazing map in terms of spacial coverage, it doesn't yet have much temporal data.
To look at glacier retreat, we will look at the USGS time series of the named glaciers of Glacier National Park2 . These 4 datasets have the contour lines of 39 glaciers for the years 1966, 1998, 2005, and 2015.
We could load and clean these datasets one by one. Copying and pasting code however is inefficient and error-prone. A better approach is to do this in a functional programming framework: create a function which does all the data loading and cleaning, then pass each element of a vector of the paths of all 4 datasets to it using purrr::map().
"Cleaning" here consists of selecting the variables we are interested in, putting them in the same order in each dataset (they were not initially) and giving the exact same name across all datasets (there were case inconsistencies between datasets and R is case sensitive).
## create a function that reads and cleans the data
prep <- function(dir) {
g <- st_read(dir)
g %<>% rename_with(~ tolower(gsub("Area....", "area", .x)))
g %<>% select(
year,
objectid,
glacname,
area,
shape_leng,
x_coord,
y_coord,
source_sca,
source
)
}
## create a vector of dataset names
dirs <- grep("GNPglaciers_.*", list.dirs(), value = T)
## pass each element of that vector through prep() thanks to map()
gnp <- map(dirs, prep)
map() returns a list, so we now have a list (gnp) of 4 elements: the 4 sf objects containing our cleaned datasets. A list is not really convenient and we will turn it into a single sf object.
Before doing so however, we want to make sure that they all have the same CRS:
st_crs(gnp[[1]]) == st_crs(gnp[[2]])
st_crs(gnp[[1]]) == st_crs(gnp[[3]])
st_crs(gnp[[1]]) == st_crs(gnp[[4]])
They do, so we can turn gnp into a single sf object:
gnp <- do.call("rbind", gnp)
gnp
str(gnp)We can now map the data:
tm_shape(gnp) +
tm_polygons("year", palette = "Blues") +
tm_layout(
title = "Glaciers of Glacier National Park",
title.position = c("center", "top"),
legend.title.color = "#fcfcfc",
legend.text.size = 1,
bg.color = "#fcfcfc",
inner.margins = c(0.07, 0.03, 0.07, 0.03),
outer.margins = 0
) +
tm_compass(
type = "arrow",
position = c("right", "top"),
text.size = 0.7
) +
tm_scale_bar(
breaks = c(0, 10, 20),
position = c("right", "BOTTOM"),
text.size = 1
)Note: I didn't want to show the legend title and because there is no option to remove it, I set its color to that of the background.

Mapping a subset of the data
Each glacier has 4 borders: one for each year of survey. They are however quite hard to see on such a large map.
Let's zoom on the Agassiz glacier:
## select the data points corresponding to the Agassiz Glacier
ag <- g %>% filter(glacname == "Agassiz Glacier")And map it:
tm_shape(ag) +
tm_polygons("year", palette = "Blues") +
tm_layout(
title = "Agassiz Glacier",
title.position = c("center", "top"),
legend.position = c("left", "bottom"),
legend.title.color = "#fcfcfc",
legend.text.size = 1,
bg.color = "#fcfcfc",
inner.margins = c(0.07, 0.03, 0.07, 0.03),
outer.margins = 0
) +
tm_compass(
type = "arrow",
position = c("right", "top"),
text.size = 0.7
) +
tm_scale_bar(
breaks = c(0, 0.5, 1),
position = c("right", "BOTTOM"),
text.size = 1
)
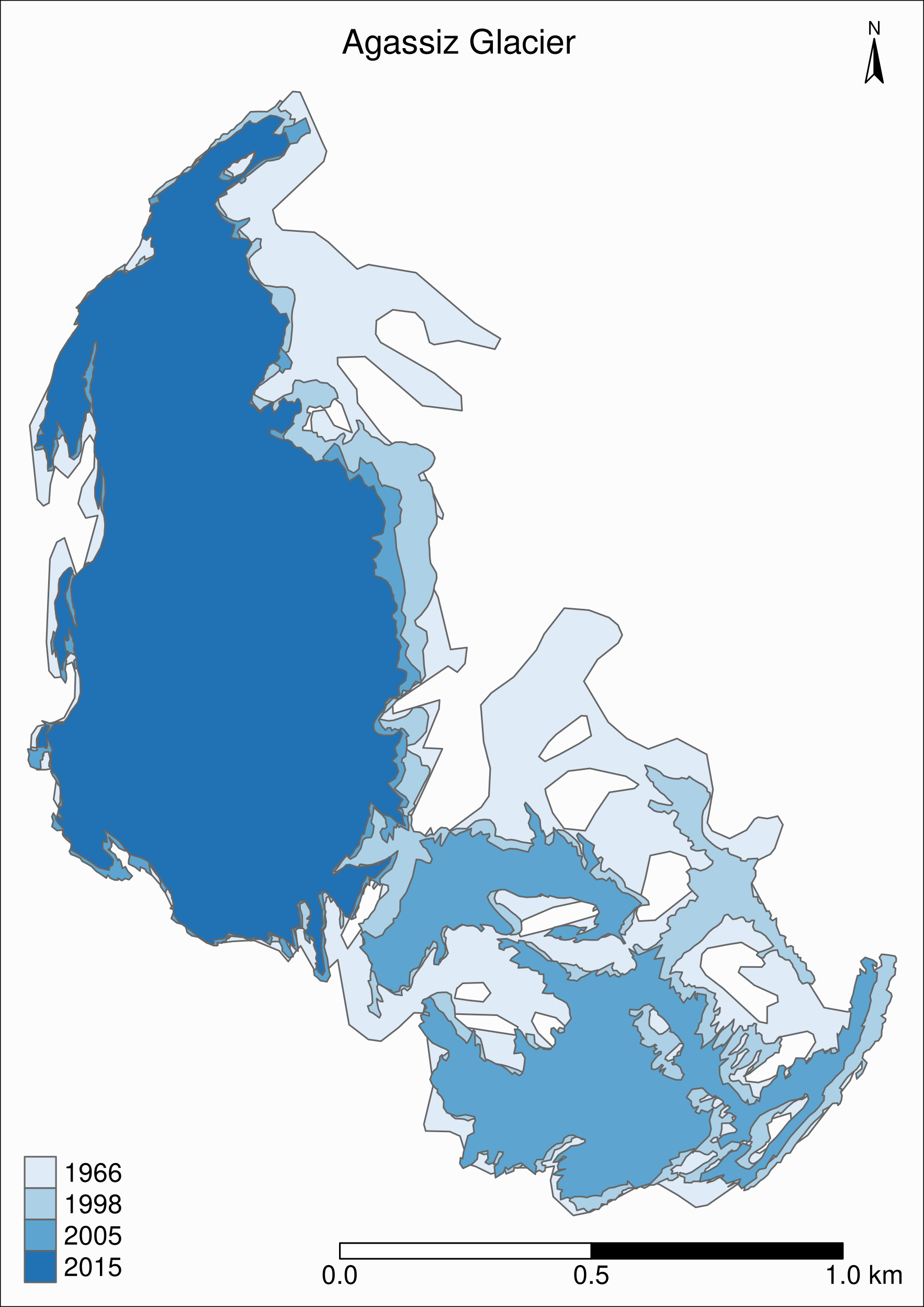
Now we can clearly see the retreat of the Agassiz Glacier between 1966 and 2015.
Additional resources
Webinar
Our next webinar on Advanced GIS mapping in R
Open GIS data:
Free GIS Data: list of free GIS datasets
Books
Geocomputation with R by Robin Lovelace, Jakub Nowosad, and Jannes Muenchow
Spatial Data Science by Edzer Pebesma, Roger Bivand
Spatial Data Science with R by Robert J. Hijmans
Using Spatial Data with R by Claudia A. Engel
Tutorial
An Introduction to Spatial Data Analysis and Visualisation in R by the CDRC
Website
r-spatial by Edzer Pebesma, Marius Appel, and Daniel Nüst
CRAN package list
Analysis of Spatial Data
Mailing list
R Special Interest Group on using Geographical data and Mapping